Classifying natural products from plants, fungi or bacteria using the COCONUT database and machine learning
Our paper Classifying natural products from plants, fungi or bacteria using the COCONUT database and machine learning is now published in Journal of Cheminformatics! The work is about distinguishing between natural products from plants, fungi, and bacteria using COCONUT, machine learning, and chemical space mapping.
Abstract:
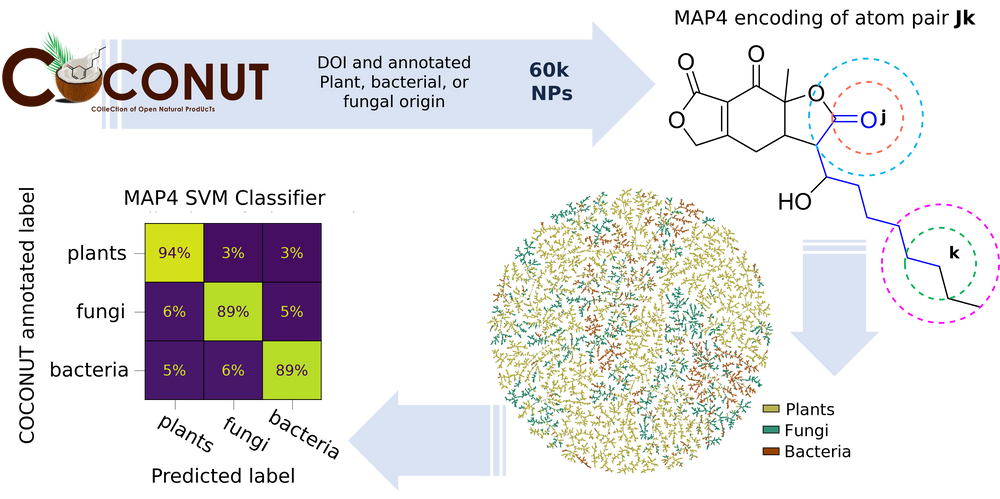
Natural products (NPs) represent one of the most important resources for discovering new drugs. Here we asked whether NP origin can be assigned from their molecular structure in a subset of 60,171 NPs in the recently reported Collection of Open Natural Products (COCONUT) database assigned to plants, fungi, or bacteria. Visualizing this subset in an interactive tree-map (TMAP) calculated using MAP4 (MinHashed atom pair fingerprint) clustered NPs according to their assigned origin (https://tm.gdb.tools/map4/coconut_tmap/), and a support vector machine (SVM) trained with MAP4 correctly assigned the origin for 94% of plant, 89% of fungal, and 89% of bacterial NPs in this subset. An online tool based on an SVM trained with the entire subset correctly assigned the origin of further NPs with similar performance (https://np-svm-map4.gdb.tools/). Origin information might be useful when searching for biosynthetic genes of NPs isolated from plants but produced by endophytic microorganisms.
Keywords:
Natural Products, Cheminformatics, Chemical Space, Visualization, Molecular Fingerprints, Machine Learning, Support Vector Machine
Author(s): Alice Capecchi, Jean-Louis Reymond