Populating Chemical Space with Peptides Using a Genetic Algorithm
The paper Populating Chemical Space with Peptides Using a Genetic Algorithm has been published in the Journal of Chemical Information and Modeling
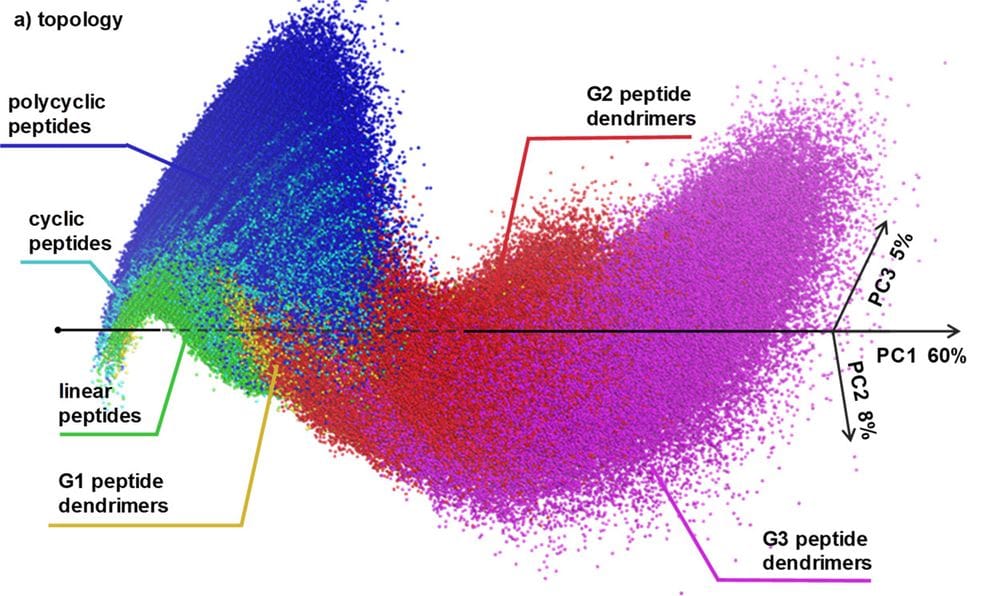
In drug discovery, one uses chemical space as a concept to organize molecules according to their structures and properties. One often would like to generate new possible molecules at a specific location in the chemical space marked by a molecule of interest. Herein, we report the peptide design genetic algorithm (PDGA, code available at https://github.com/reymond-group/PeptideDesignGA), a computational tool capable of producing peptide sequences of various topologies (linear, cyclic/polycyclic, or dendritic) in proximity of any molecule of interest in a chemical space defined by macromolecule extended atom-pair fingerprint (MXFP), an atom-pair fingerprint describing molecular shape and pharmacophores. We show that the PDGA generates high-similarity analogues of bioactive peptides with diverse peptide chain topologies and of nonpeptide target molecules. We illustrate the chemical space accessible by the PDGA with an interactive 3D map of the MXFP property space available at http://faerun.gdb.tools/. The PDGA should be generally useful to generate peptides at any location in the chemical space.
Author(s): Alice Capecchi, Alain Zhang, Jean-Louis Reymond